KRISP NEWS


Representatives from the Mastercard Foundation visited CERI to meet African STARS fellows, learn about their experiences, and explore how the programme is helping develop the next generation of African researchers, innovators, and public health leaders.
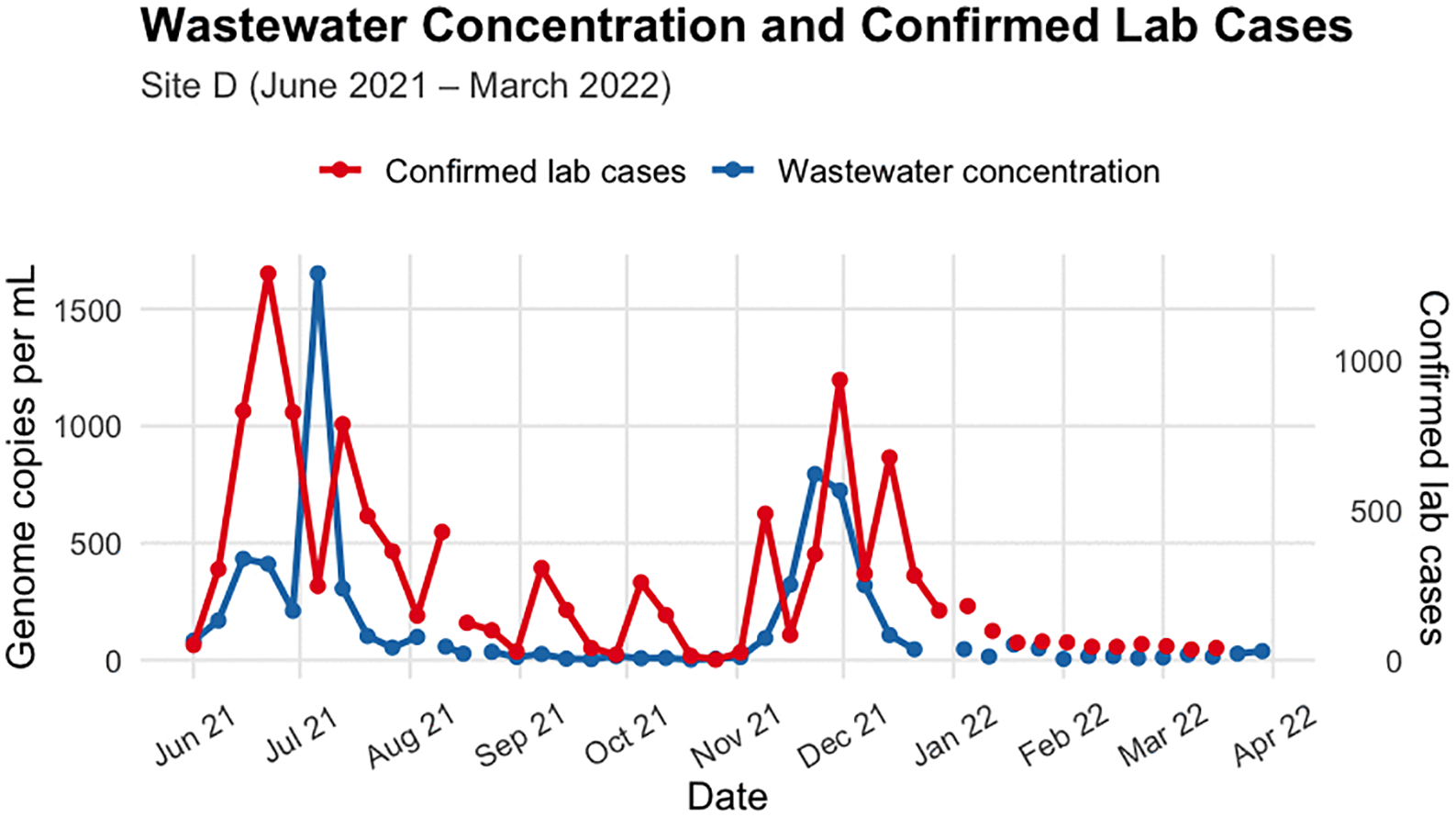
A new study highlights the potential of wastewater surveillance as a cost-effective tool for tracking disease outbreaks in low-resource settings. By comparing wastewater data with clinical testing results during a SARS-CoV-2 outbreak in South Africa, researchers found that wastewater monitoring can provide valuable early insights into community-level disease trends.

An international symposium explored how research on oncogenic viruses is advancing cancer prevention, diagnosis, and treatment, with a focus on translating scientific discoveries into public health impact in Africa.

Behind every successful research programme is a strong operational foundation. At CERI, Zethu Luthulis dedication to research management and administration helps create the conditions that enable researchers and students to thrive.

Researchers and community engagement teams from four African countries gathered in Kenya for the 2026 WEMA Consortium and Scientific Advisory Board meeting, sharing emerging findings on climate-related mental health and shaping the next phase of the project.

The Wastewater & Environmental Surveillance Meeting 2026 in Ghana brought together researchers from across Africa to explore how wastewater data can support disease surveillance, outbreak preparedness, and public health decision-making.
KRISP SCIENTIFIC PUBLICATIONS
Addressing pandemic-wide systematic errors in the SARS-CoV-2 phylogeny.
Hunt M, Hinrichs A, Anderson D, Karim L, Dearlove B, Knaggs J, Constantinides B, Fowler P, Rodger G, Street T, Lumley S, Webster H, Sanderson T, Ruis C, Kotzen B, de Maio N, Amenga-Etego L, Amuzu D, Avaro M, Awandare G, Ayivor-Djanie R, Barkham T, Bashton M, Batty E, Bediako Y, De Belder D, Benedetti E, Bergthaler A, Boers S, Campos J, Carr R, Chen Y, Cuba F, Dattero M, Dejnirattisai W, Dilthey A, Duedu K, Endler L, Engelmann I, Francisco N, Fuchs J, Gnimpieba E, Groc S, Gyamfi J, Heemskerk D, Houwaart T, Hsiao N, Huska M, Hölzer M, Iranzadeh A, Jarva H, Jeewandara C, Jolly B, Joseph R, Kant R, Ki K, Kurkela S, Lappalainen M, Lataretu M, Lemieux J, Liu C, Malavige G, Mashe T, Mongkolsapaya J, Montes B, Mora J, Moranga C, Mvula B, Nagarajan N, Nelson A, Ngoi J, da Paixão J, Panning M, Poklepovich T, Quashie P, Ranasinghe D, Russo M, San J, Sanderson N, Scaria V, Screaton G, Sessions O, Sironen T, Sisay A, Smith D, Smura T, Supasa P, Suphavilai C, Swann J, Tegally H, Tegomoh B, Vapalahti O, Walker A, Wilkinson R, Williamson C, Zair X, Biere B, Dürrwald R, Mache C, Oh D, Schulze J, Wedde M, Wolff T, Fuchs S, Semmler T, Paraskevopoulou S, Kerber R, Kröger S, Haas W, Bode K, Corman V, Erren M, Finzer P, Grosser R, Haffner M, Hermann B, Kiel C, Krumbholz A, Lorentz T, Meinck K, Nitsche A, Petzold M, Schwanz T, Szabados F, Tewald F, Tiemann C, de Oliveira T, Peto T, Crook D, Corbett-Detig R, Iqbal Z, Nature Methods (2026), 23:653-662.
Tracing the spatial origins and spread of SARS-CoV-2 Omicron lineages in South Africa.
Dor G, Wilkinson E, Martin DP, Moir M, Tshiabuila D, Kekana D, Ntozini B, Joseph R, Iranzadeh A, Nyaga MM, Goedhals D, Maponga T, Maritz J, Laguda-Akingba O, Ramphal Y, MacIntyre C, Chabuka L, Pillay S, Giandhari J, Baxter C, Hsiao NY, Preiser W, Bhiman JN, Davies MA, Venter M, Treurnicht FK, Wolter N, Williamson C, von Gottberg A, Lessells R, Tegally H, de Oliveira T, Nature Communications (2025), 28;16(1):4937. doi: 10.1038/s41467-025-60081-0:.
Genomic Surveillance of Climate-Amplified Cholera Outbreak, Malawi, 20222023.
Chabuka L, Choga W, Mavian C, Moir M, Morgenstern C, Tegaly H, Sharma A, Wilkinson E, Naidoo Y, Inward R, Bhatt S, WilliamWint G, Khan K, Bogoch I, Kraemer M, Lourenço J, Baxter C, Tagliamonte M, Salemi M, Lessells R, Mitambo C, Chitatanga R, Bitilinyu-Bango J, Chiwaula M, Chavula Y, Bukhu M, Manda H, Chitenje M, Malolo I, Mwanyongo A, Mvula B, Nyenje M, de Oliveira T, Kagoli M, Emerging Infectious Diseases (2025), 31(6):. doi: 10.3201/eid3106.240930.:.
Importance of outbreak response research in bridging knowledge gaps on emerging infectious diseases.
Breiman R, Osoro E, Reithinger R, Wang D, Diamond M, Van Voorhis W, Wasserheit J, Rabinowitz P, Mboup S, Hemingway-Foday J, de Oliveira T, Boon A, Schieffelin J, Sempowski G, Moody M, Vasilakis N, Hanley K, Nasimiyu C, Situma S, Ngere I, Kyobe Bosa H, Nyakarahuka L, Bakamutumaho B, Woodson S, Njenga M, BMJ Global Health (2025), 10(6):e018297. doi: 10.1136/bmjgh-2024-018297.:.
Artificial intelligence for modelling infectious disease epidemics.
Kraemer M, Tsui J, Chang S, Lytras S, Khurana M, Vanderslott S, Bajaj S, Scheidwasser N, Curran-Sebastian J, Semenova E, Zhang M, Unwin H, Watson O, Mills C, Dasgupta A, Ferretti L, Scarpino S, Koua E, Morgan O, Tegally H, Paquet U, Moutsianas L, Fraser C, Ferguson N, Topol E, Duchêne D, Stadler T, Kingori P, Parker M, Dominici F, Shadbolt N, Suchard M, Ratmann O, Flaxman S, Holmes E, Gomez-Rodriguez M, Schölkopf B, Donnelly C, Pybus O, Cauchemez S, Bhatt S, Nature (2025), :.
Spatiotemporal disease suitability prediction for Oropouche virus and the role of vectors across the Americas.
Poongavanan J, Dunaiski M, Dor G, Kraemer M, Giovanetti M, Lim A, Brady O, Baxter C, Fonseca V, Alcantara L, de Oliveira T, Tegally H, medRxiv (2025), doi: 10.1101/2025.02.28.25323068.:.
Characterization of SARS-CoV-2 intrahost genetic evolution in vaccinated and non-vaccinated patients from the Kenyan population.
Lugano D, Mwangi K, Mware B, Kibet G, Osiany S, Kiritu E, Dobi P, Muli C, Njeru R, de Oliveira T, Njenga M, Routh A, Oyola S, medRxiv (2025), doi: 10.1101/2025.03.03.25323296.:.
KRISP VIDEOS
COVID-19 | News sub-variant being monitored closely
By: Tulio De Oliveira and CERI and KRISP teams
KRISP BIOINFORMATICS TOOLS
Genome Detective Coronavirus Typing Tool
Genome Detective Coronavirus Typing Tool for rapid identification and characterization of novel coronavirus genomes
Genome Detective Dengue Virus Typing Tool
This is a beta version of our Dengue Virus Typing tool. For the mean time, this tool should be used for evaluation only. Please send feedback to Tulio de Oliveira.
Genome Detective Zika Typing Tool
This is the first version of the Zika typing tool, which uses phylogenetic analysis to identify the species and genotype of the virus.
Genome Detective Chikungunya Typing Tool
This is the first version of the Chikungunya typing tool, which uses phylogenetic analysis to identify the species and genotype of the virus.
Genome Detective Yellow Fever Virus Typing Tool
This is the first version of the Yellow Fever typing tool, which uses phylogenetic analysis to identify the species and genotype of the virus.
This is the first version of our Arbovirus typing tool for Chikungunya, Dengue, Yellow Fever and Zika
REGA HIV Subtyping Tool V3 - Belgium Mirror
Phylogenetic tool to identify the HIV-1 subtypes and recombinants. Query sequences are analysed for recombination using bootscanning methods. The version 3 contains new CRFs (CRF01_AE to CRF47_BF).
MORE TOOLS
KRISP has been created by the coordinated effort of the University of KwaZulu-Natal (UKZN), the Technology Innovation Agency (TIA) and the South African Medical Research Countil (SAMRC).
Location: K-RITH Tower Building
Nelson R Mandela School of Medicine, UKZN
719 Umbilo Road, Durban, South Africa.
Director: Prof. Tulio de Oliveira