KRISP NEWS

A connection made at SWEAT Africa 2026 led to a role at Nanosene within weeks, offering a clear example of how the event creates the conditions for real opportunities to emerge.
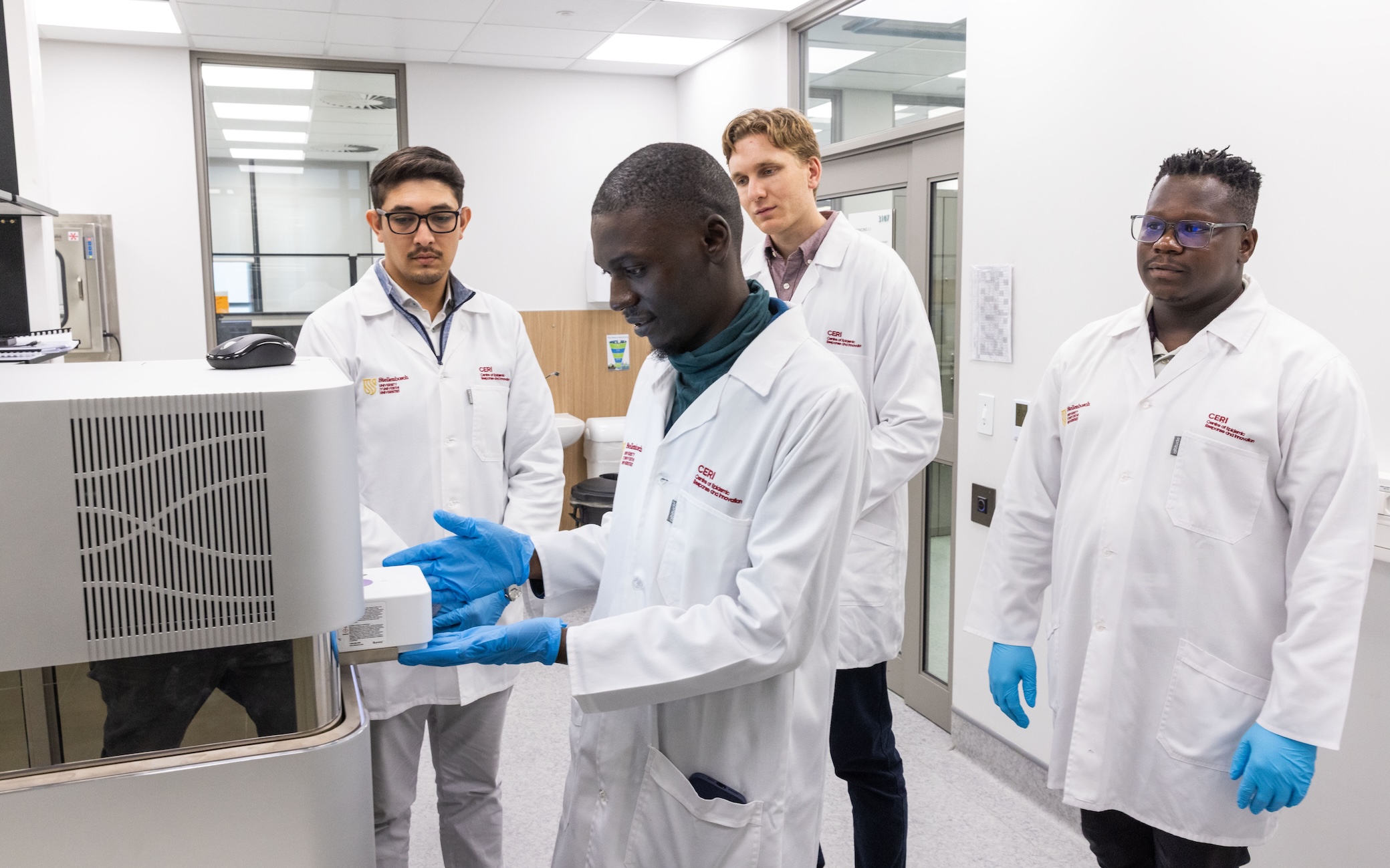

As emerging and re-emerging viruses continue to shape global public health, the ability to rapidly detect and characterise pathogens has become increasingly important, particularly in regions where surveillance resources remain unevenly distributed. Earlier this month, scientists, laboratory professionals, and public health researchers from across Africa gathered at the Centre for Epidemic Response and Innovation (CERI) at Stellenbosch University for an intensive hands-on workshop focused on one of the fields most advanced viral sequencing approaches: VirCapSeq-VERT.

Selected from more than 900 international applicants as one of only 10 young innovators invited to the United Nations Science, Technology and Innovation Forum in New York, African STARS fellow Kennedy Mulungu is emerging as part of a new generation of African scientists and entrepreneurs building globally competitive innovation from the continent.

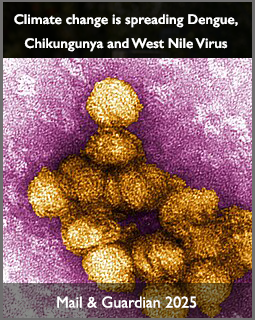
As climate and environmental change continue to reshape ecosystems across the world, scientists are warning that mosquito-borne diseases may spread into new regions, become more frequent, and intensify in areas already under pressure. A major new international research grant awarded to Dr Monika Moir at the Centre for Epidemic Response and Innovation (CERI) will place Africa at the centre of efforts to better understand one of these moving threats: West Nile virus.

Prof Tulio de Oliveira, Director of the Centre for Epidemic Response and Innovation (CERI) at Stellenbosch University and founder and director of the KwaZulu-Natal Research Innovation and Sequencing Platform (KRISP) at the University of KwaZulu-Natal, has been awarded the Order of Mapungubwe in Gold by President Cyril Ramaphosa, in recognition of more than two decades of work building South Africa's capacity to detect, track, and respond to infectious disease threats.

A SARS-CoV-2 lineage once thought extinct has re-emerged after more than two years revealing how the virus can evolve undetected and return with significant genetic change.

Collaboration and continuous learning are at the heart of advancing genomics in Africa and the return of Dr Armando Djiyou to KRISP reflects just that.
KRISP SCIENTIFIC PUBLICATIONS
Addressing pandemic-wide systematic errors in the SARS-CoV-2 phylogeny.
Hunt M, Hinrichs A, Anderson D, Karim L, Dearlove B, Knaggs J, Constantinides B, Fowler P, Rodger G, Street T, Lumley S, Webster H, Sanderson T, Ruis C, Kotzen B, de Maio N, Amenga-Etego L, Amuzu D, Avaro M, Awandare G, Ayivor-Djanie R, Barkham T, Bashton M, Batty E, Bediako Y, De Belder D, Benedetti E, Bergthaler A, Boers S, Campos J, Carr R, Chen Y, Cuba F, Dattero M, Dejnirattisai W, Dilthey A, Duedu K, Endler L, Engelmann I, Francisco N, Fuchs J, Gnimpieba E, Groc S, Gyamfi J, Heemskerk D, Houwaart T, Hsiao N, Huska M, Hölzer M, Iranzadeh A, Jarva H, Jeewandara C, Jolly B, Joseph R, Kant R, Ki K, Kurkela S, Lappalainen M, Lataretu M, Lemieux J, Liu C, Malavige G, Mashe T, Mongkolsapaya J, Montes B, Mora J, Moranga C, Mvula B, Nagarajan N, Nelson A, Ngoi J, da Paixão J, Panning M, Poklepovich T, Quashie P, Ranasinghe D, Russo M, San J, Sanderson N, Scaria V, Screaton G, Sessions O, Sironen T, Sisay A, Smith D, Smura T, Supasa P, Suphavilai C, Swann J, Tegally H, Tegomoh B, Vapalahti O, Walker A, Wilkinson R, Williamson C, Zair X, Biere B, Dürrwald R, Mache C, Oh D, Schulze J, Wedde M, Wolff T, Fuchs S, Semmler T, Paraskevopoulou S, Kerber R, Kröger S, Haas W, Bode K, Corman V, Erren M, Finzer P, Grosser R, Haffner M, Hermann B, Kiel C, Krumbholz A, Lorentz T, Meinck K, Nitsche A, Petzold M, Schwanz T, Szabados F, Tewald F, Tiemann C, de Oliveira T, Peto T, Crook D, Corbett-Detig R, Iqbal Z, Nature Methods (2026), 23:653-662.
Tracing the spatial origins and spread of SARS-CoV-2 Omicron lineages in South Africa.
Dor G, Wilkinson E, Martin DP, Moir M, Tshiabuila D, Kekana D, Ntozini B, Joseph R, Iranzadeh A, Nyaga MM, Goedhals D, Maponga T, Maritz J, Laguda-Akingba O, Ramphal Y, MacIntyre C, Chabuka L, Pillay S, Giandhari J, Baxter C, Hsiao NY, Preiser W, Bhiman JN, Davies MA, Venter M, Treurnicht FK, Wolter N, Williamson C, von Gottberg A, Lessells R, Tegally H, de Oliveira T, Nature Communications (2025), 28;16(1):4937. doi: 10.1038/s41467-025-60081-0:.
Genomic Surveillance of Climate-Amplified Cholera Outbreak, Malawi, 20222023.
Chabuka L, Choga W, Mavian C, Moir M, Morgenstern C, Tegaly H, Sharma A, Wilkinson E, Naidoo Y, Inward R, Bhatt S, WilliamWint G, Khan K, Bogoch I, Kraemer M, Lourenço J, Baxter C, Tagliamonte M, Salemi M, Lessells R, Mitambo C, Chitatanga R, Bitilinyu-Bango J, Chiwaula M, Chavula Y, Bukhu M, Manda H, Chitenje M, Malolo I, Mwanyongo A, Mvula B, Nyenje M, de Oliveira T, Kagoli M, Emerging Infectious Diseases (2025), 31(6):. doi: 10.3201/eid3106.240930.:.
Importance of outbreak response research in bridging knowledge gaps on emerging infectious diseases.
Breiman R, Osoro E, Reithinger R, Wang D, Diamond M, Van Voorhis W, Wasserheit J, Rabinowitz P, Mboup S, Hemingway-Foday J, de Oliveira T, Boon A, Schieffelin J, Sempowski G, Moody M, Vasilakis N, Hanley K, Nasimiyu C, Situma S, Ngere I, Kyobe Bosa H, Nyakarahuka L, Bakamutumaho B, Woodson S, Njenga M, BMJ Global Health (2025), 10(6):e018297. doi: 10.1136/bmjgh-2024-018297.:.
Artificial intelligence for modelling infectious disease epidemics.
Kraemer M, Tsui J, Chang S, Lytras S, Khurana M, Vanderslott S, Bajaj S, Scheidwasser N, Curran-Sebastian J, Semenova E, Zhang M, Unwin H, Watson O, Mills C, Dasgupta A, Ferretti L, Scarpino S, Koua E, Morgan O, Tegally H, Paquet U, Moutsianas L, Fraser C, Ferguson N, Topol E, Duchêne D, Stadler T, Kingori P, Parker M, Dominici F, Shadbolt N, Suchard M, Ratmann O, Flaxman S, Holmes E, Gomez-Rodriguez M, Schölkopf B, Donnelly C, Pybus O, Cauchemez S, Bhatt S, Nature (2025), :.
Spatiotemporal disease suitability prediction for Oropouche virus and the role of vectors across the Americas.
Poongavanan J, Dunaiski M, Dor G, Kraemer M, Giovanetti M, Lim A, Brady O, Baxter C, Fonseca V, Alcantara L, de Oliveira T, Tegally H, medRxiv (2025), doi: 10.1101/2025.02.28.25323068.:.
Characterization of SARS-CoV-2 intrahost genetic evolution in vaccinated and non-vaccinated patients from the Kenyan population.
Lugano D, Mwangi K, Mware B, Kibet G, Osiany S, Kiritu E, Dobi P, Muli C, Njeru R, de Oliveira T, Njenga M, Routh A, Oyola S, medRxiv (2025), doi: 10.1101/2025.03.03.25323296.:.
KRISP VIDEOS
COVID-19 | News sub-variant being monitored closely
By: Tulio De Oliveira and CERI and KRISP teams
KRISP BIOINFORMATICS TOOLS
Genome Detective Coronavirus Typing Tool
Genome Detective Coronavirus Typing Tool for rapid identification and characterization of novel coronavirus genomes
Genome Detective Dengue Virus Typing Tool
This is a beta version of our Dengue Virus Typing tool. For the mean time, this tool should be used for evaluation only. Please send feedback to Tulio de Oliveira.
Genome Detective Zika Typing Tool
This is the first version of the Zika typing tool, which uses phylogenetic analysis to identify the species and genotype of the virus.
Genome Detective Chikungunya Typing Tool
This is the first version of the Chikungunya typing tool, which uses phylogenetic analysis to identify the species and genotype of the virus.
Genome Detective Yellow Fever Virus Typing Tool
This is the first version of the Yellow Fever typing tool, which uses phylogenetic analysis to identify the species and genotype of the virus.
This is the first version of our Arbovirus typing tool for Chikungunya, Dengue, Yellow Fever and Zika
REGA HIV Subtyping Tool V3 - Belgium Mirror
Phylogenetic tool to identify the HIV-1 subtypes and recombinants. Query sequences are analysed for recombination using bootscanning methods. The version 3 contains new CRFs (CRF01_AE to CRF47_BF).
MORE TOOLS
KRISP has been created by the coordinated effort of the University of KwaZulu-Natal (UKZN), the Technology Innovation Agency (TIA) and the South African Medical Research Countil (SAMRC).
Location: K-RITH Tower Building
Nelson R Mandela School of Medicine, UKZN
719 Umbilo Road, Durban, South Africa.
Director: Prof. Tulio de Oliveira